library(ggalign)
#> Loading required package: ggplot2
#> ========================================
#> ggalign version 1.2.0.9000
#>
#> If you use it in published research, please cite:
#> Peng, Y.; Jiang, S.; Song, Y.; et al. ggalign: Bridging the Grammar of Graphics and Biological Multilayered Complexity. Advanced Science. 2025. doi:10.1002/advs.202507799
#> ========================================Visualizing Phylogenetic Trees
ggalign provides direct support for phylo objects from the ape package, allowing you to visualize phylogenetic trees alongside other data in an integrated way.
Note that you can also use stack_discrete() to display the tree in a stacked arrangement rather than in a radial layout. For this documentation, we illustrate examples using circle_discrete() as it is the most commonly used approach.
Basic phylogenetic tree alignment
circle_discrete(x,
radial = coord_radial(inner.radius = 0.2, rotate.angle = TRUE)
) +
align_phylo(tr) +
ggalign() +
geom_tile(aes(y = .column_index, fill = value)) +
scale_fill_viridis_c()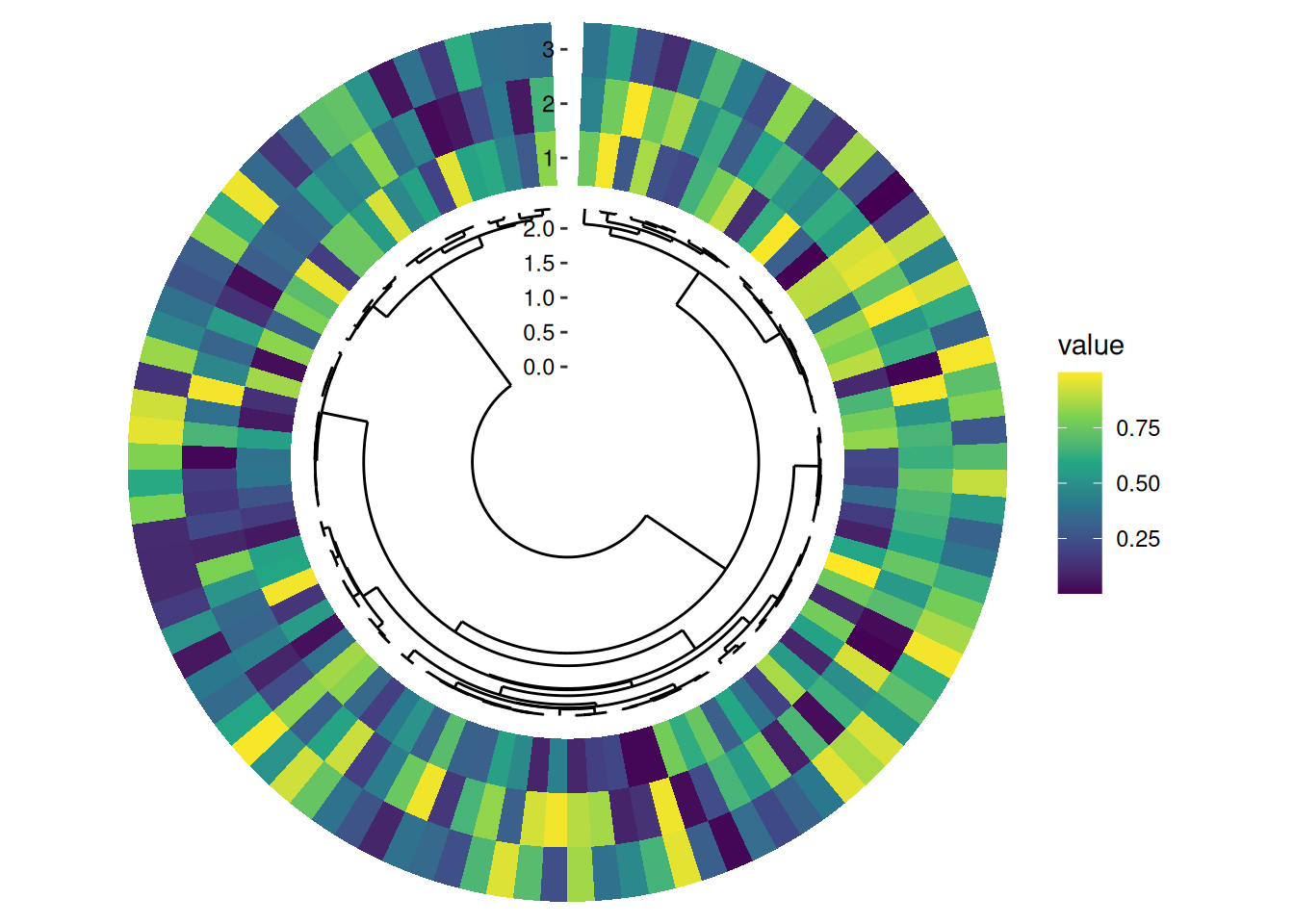
The underlying data passed to align_phylo() includes both tip and internal node information. Edge data is stored as a ggalign attribute, and you can retrieve it using ggalign_attr(). See ?fortify_data_frame.phylo for details.
circle_discrete(x,
radial = coord_radial(inner.radius = 0.2, rotate.angle = TRUE)
) +
align_phylo(tr) +
geom_point() +
ggalign() +
geom_tile(aes(y = .column_index, fill = value)) +
scale_fill_viridis_c() &
scale_y_continuous(breaks = NULL)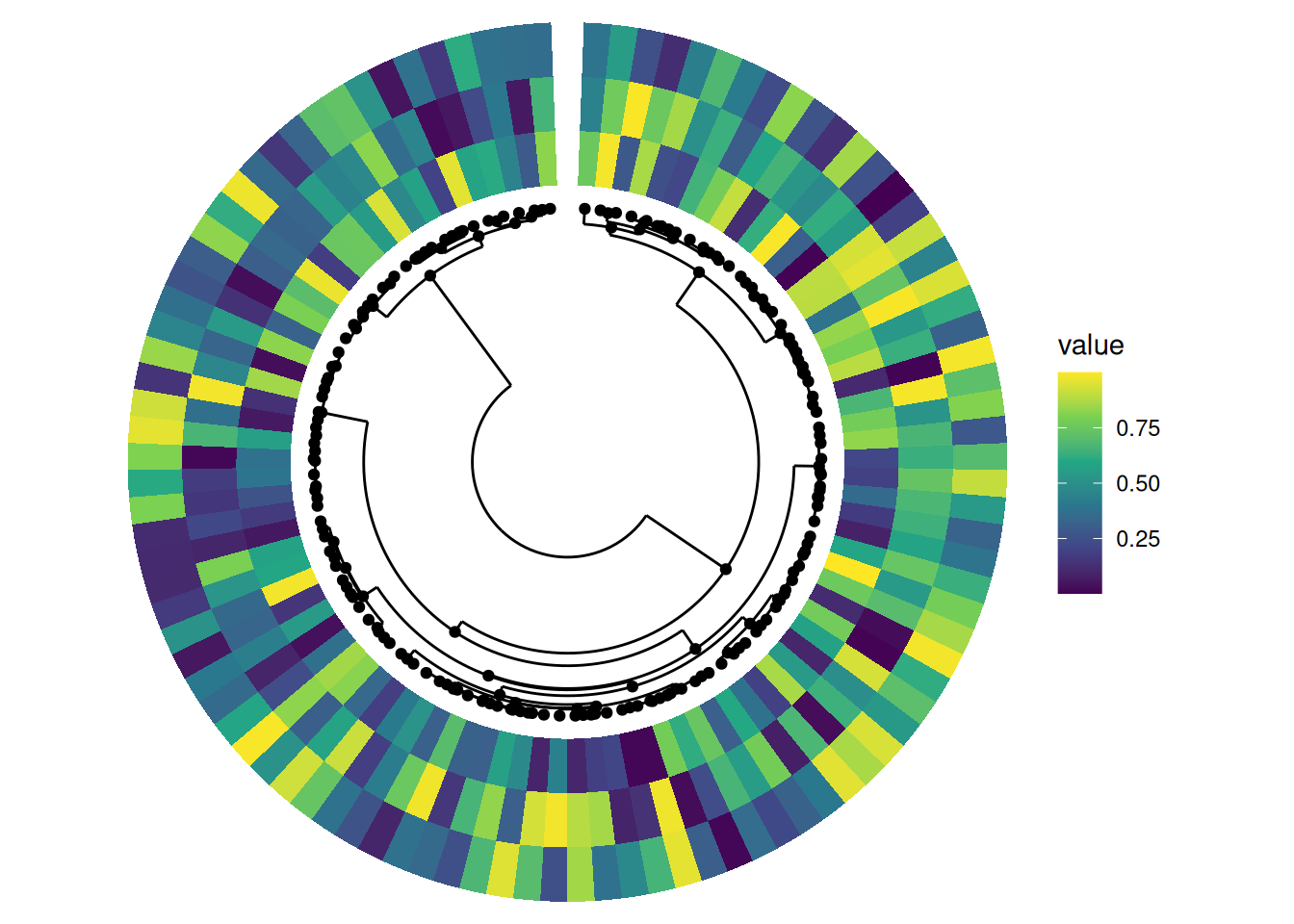
Grouping by clades
By default, align_phylo() will add a single phylogenetic tree, and align_group() can then facet the data by clade:
circle_discrete(x,
radial = coord_radial(inner.radius = 0.2, rotate.angle = TRUE)
) +
align_phylo(tr, mapping = aes(color = clade)) +
geom_point() +
align_group(groups) +
ggalign() +
geom_tile(aes(y = .column_index, fill = value)) +
scale_fill_viridis_c() &
scale_y_continuous(breaks = NULL)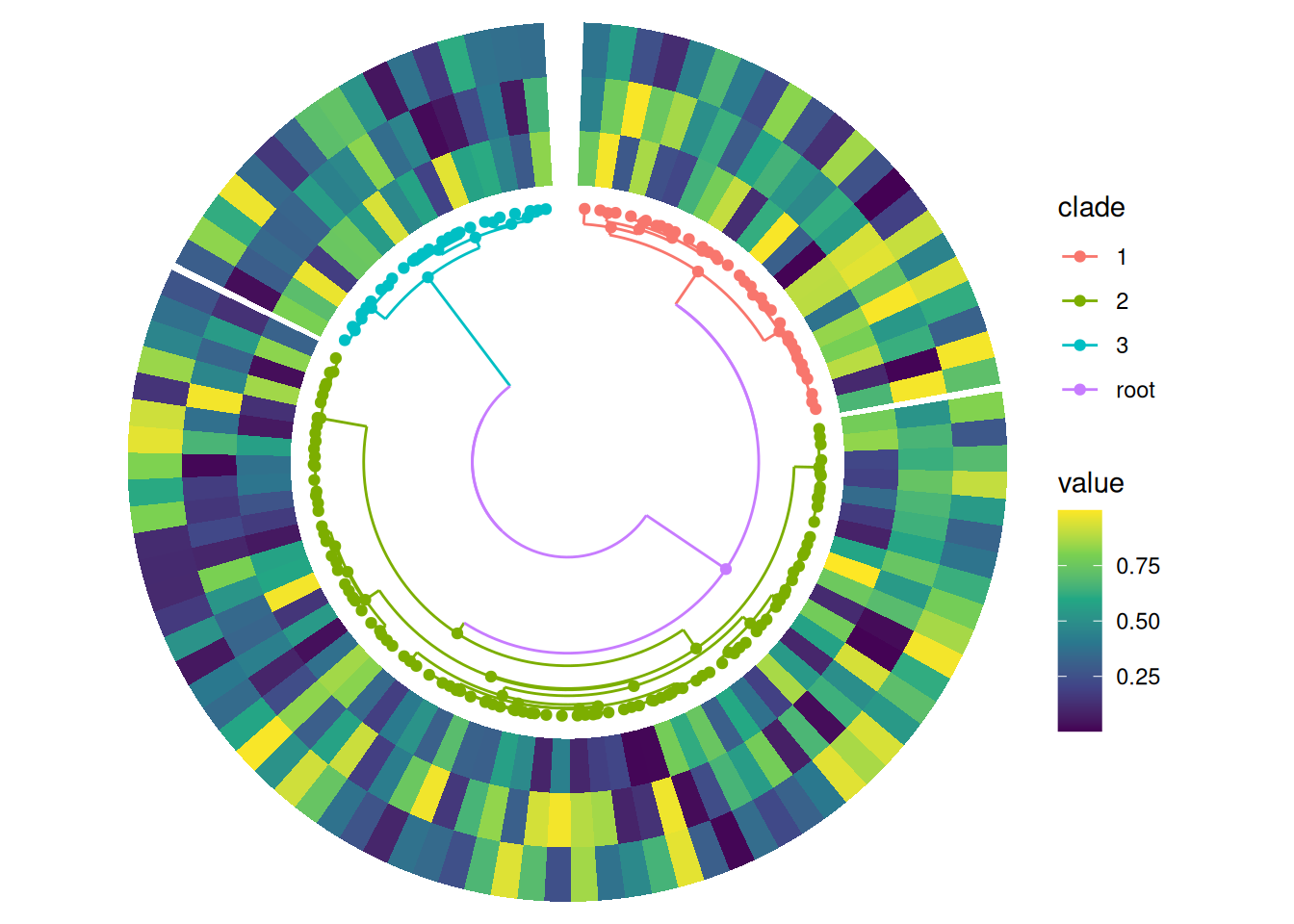
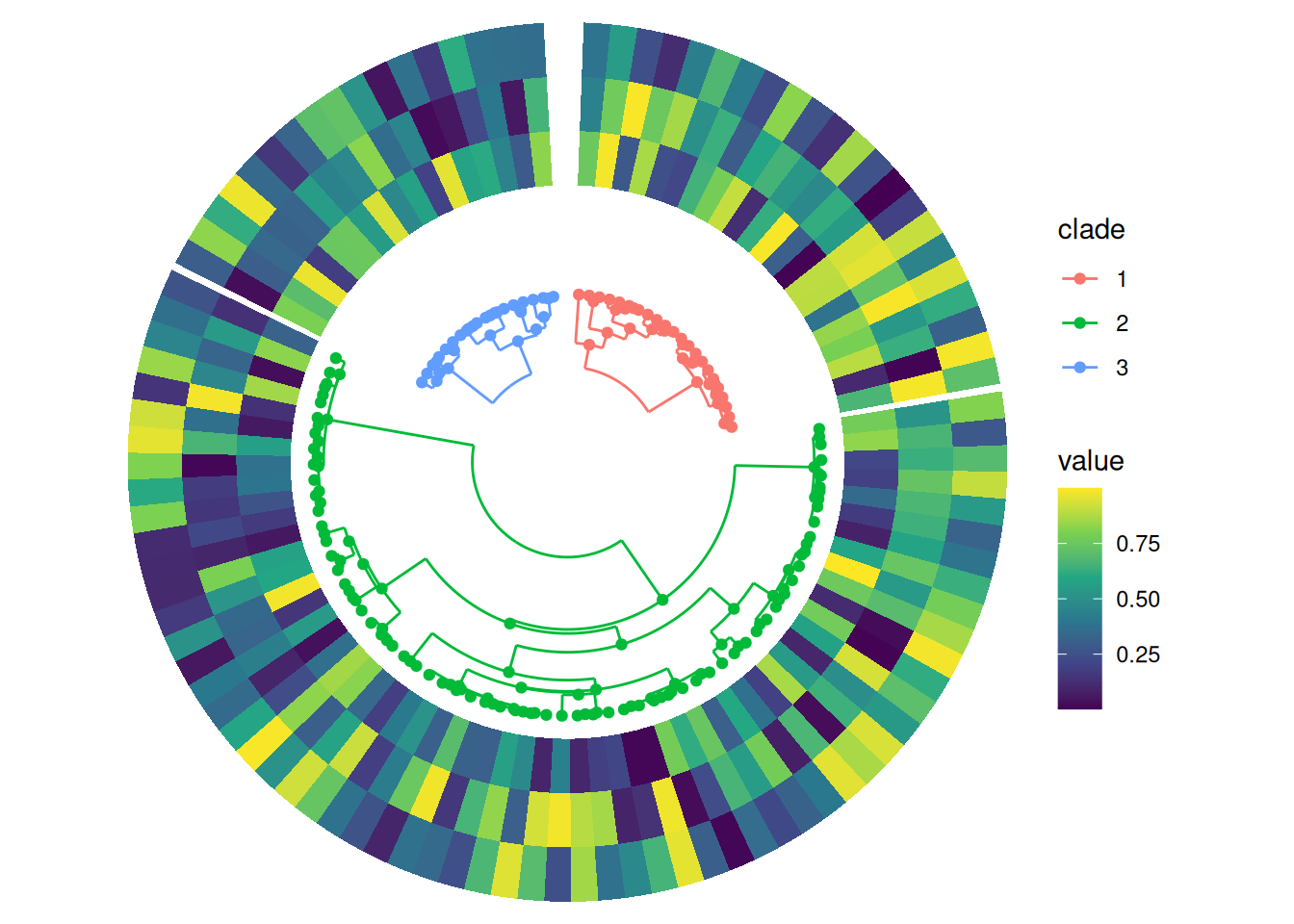
Splitting into multiple subtrees
If groups are already set before adding the phylogenetic tree, you can set split = TRUE in align_phylo() to automatically split the tree into multiple subtrees, one for each group.
circle_discrete(x,
radial = coord_radial(inner.radius = 0.2, rotate.angle = TRUE)
) +
align_group(groups) +
align_phylo(tr, mapping = aes(color = clade), split = TRUE) +
geom_point() +
ggalign() +
geom_tile(aes(y = .column_index, fill = value)) +
scale_fill_viridis_c() &
scale_y_continuous(breaks = NULL)